NewScientist.com news service
Duncan Graham-Rowe
A map of the Americas measuring just a few hundred nanometres across has been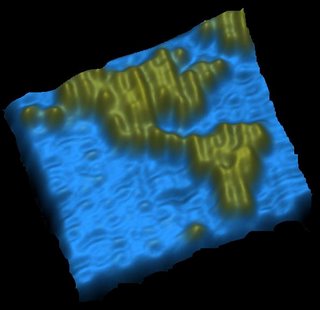 created out of meticulously folded strands of DNA, using a new technique for manipulating molecules dubbed "DNA origami".
created out of meticulously folded strands of DNA, using a new technique for manipulating molecules dubbed "DNA origami".
The nanoscale map, which sketches out both North and South America at a staggering 200-trillionths of their actual size, aims to demonstrate the precision and complexity with which DNA can be manipulated using the approach.
According to the map's creator, Paul Rothemund at Caltech in Pasadena, US, DNA origami could prove hugely important for building future nano-devices including molecular machines and quantum computer components. The technique exploits the fact that complementary base pairs of DNA will automatically stick together and involves folding a single strand of DNA in many different ways.
To make the nano-map, Rothemund first needed to create a suitable "canvas". He used a single strand of DNA from a bacteria-destroying virus called M13 mp18. The strand was folded over and over at regular intervals using smaller strands of complementary DNA, which pull t wo parts of the strand together to create each fold.
wo parts of the strand together to create each fold.
The result is a flat surface made from a long double helix, comprised of the single long strand and more than 200 shorter strands stuck along its length, "stapling" it together at key locations.
Quantum dots
If this helix was a perfectly formed, the canvas would appear blank. But by using short strands of DNA, containing bits that do not stick to the main strand, it is possible to cause parts of the canvas stand out . Precisely controlling the location of these extra bits made it possible for Rothemund to draw out the shape of the map.
To make the design process less complicated, Rothemund created software that works out which short-strand sequences will generate different shapes. Rothemund says it should be possible to precisely arrange quantum dots or carbon nanotubes after chemically binding them to DNA, using the method.
William Shih at the Biomolecular Nanotechnology Group at Harvard Medical School in Boston, US, says this offers the most flexible method yet for building nanoscale structures. Shih is experimenting with the technique as a means of making molecular 3D cages, which could be used to build molecular motors.
Disposable scaffolds
It is not the first time DNA has been used to make structures - the idea was originally developed by Nadrian Seeman at New York University, US - Rothemund's approach takes things to a new level of complexity.
Although Rothemund has only made 2D shapes there is nothing to prevent the technique
being applied to make complex 3D structures, he says. These could serve as disposable scaffolds to help molecules and carbon nanotubes self assemble.
Other examples "DNA artwork", creating using the same technique, include "smiley-faces", complex geometric shapes and a picture of a double helix with the letters "DNA" running above it.


1 comment:
huhngh.
Post a Comment